Blood cancer profiling challenges with NGS
Hematology oncology research has greatly benefited from next generation sequencing (NGS) methods to identify relevant cancer-driving biomarkers. Heme malignancies have a complex biomarker spectrum, but NGS methods to detect relative fusions, splicing, expression, copy number variants, single nucleotide variants, and complex alterations have been vital to labs.
However, traditional NGS methods may not provide all relevant information and can have difficult workflows and analysis. One specific challenge in blood cancer profiling is myeloid-relevant genes rich in GC-content, such as CEBPA, are difficult to amplify and sequence with NGS. Additionally, internal tandem duplications (ITDs), such as FLT3-ITD relevant in AML, can be difficult to detect with NGS due to variable locations and lengths along with random sequence insertions which can confound genome aligners. Due to this, most NGS methods have difficulty detecting ITDs of all sizes and insertion points, and sensitivity dramatically reduces above 100 bp ITDs. Finally, there are a multitude of relevant fusions in hematological neoplasms, including many novel fusions. In fact, one study found that 41% of acute myeloid leukemias and 88% of myelodysplastic syndromes had novel fusions [1]. However, since these are relatively rare, traditional opposing primer-based NGS methods may not be designed with primers to detect all of them, thus missing potentially relevant biomarkers (Figure 1).
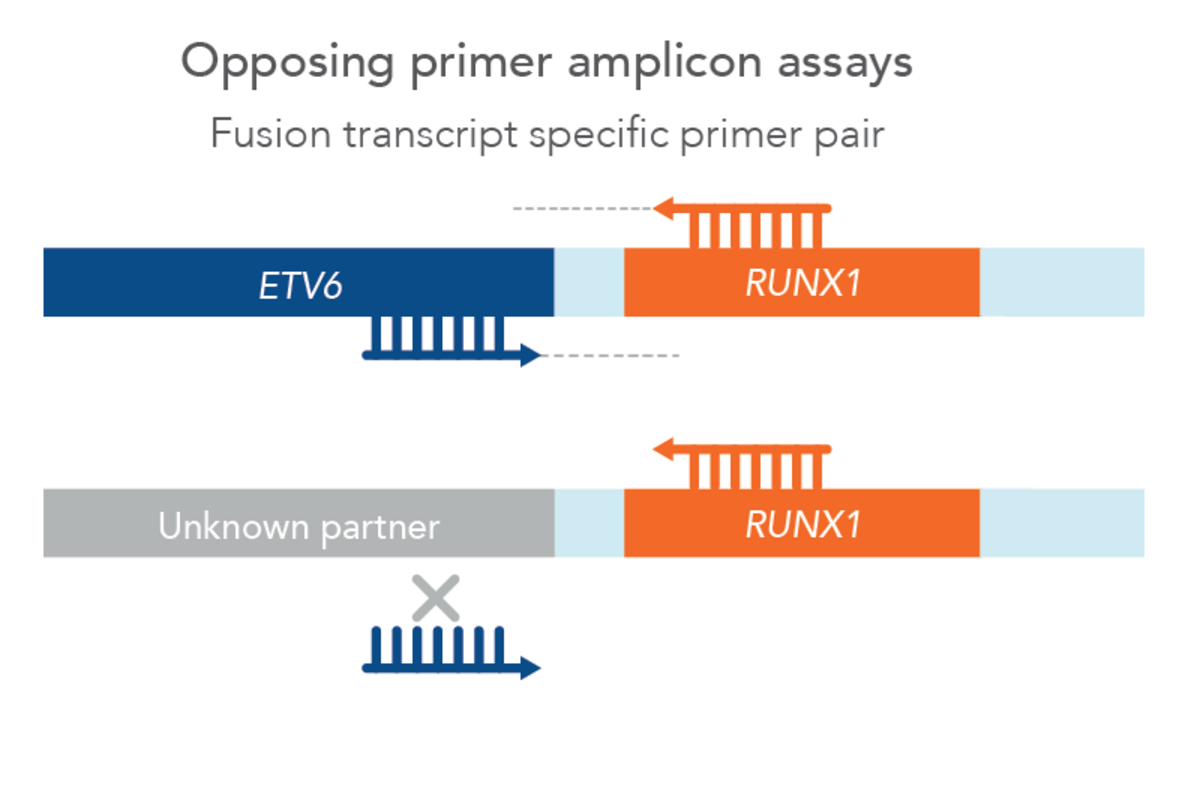
Additionally, NGS library preparation workflows can be cumbersome, increasing the opportunity for manual errors. In-house bioinformatics pipelines can be expensive and time-consuming to build and maintain. They also may lack the QC and statistical tools necessary to reduce errors and guide calling, leaving it up to lab staff to make difficult calls.
Detect more with AMP chemistry
For blood cancer profiling, Anchored Multiplex PCR (AMP) is a target enrichment method that uses molecular barcoded adapters and single, nested, gene-specific primers for amplification, permitting open-ended capture of DNA, RNA, and cfDNA fragments. This approach enables the detection of novel fusions (Figure 2), allows for flexible primer design, and protects against dropout of primers. MBC adapter ligation prior to PCR combined with Archer Analysis software power the correction of systematic errors and quantitative analysis of the original unamplified template.
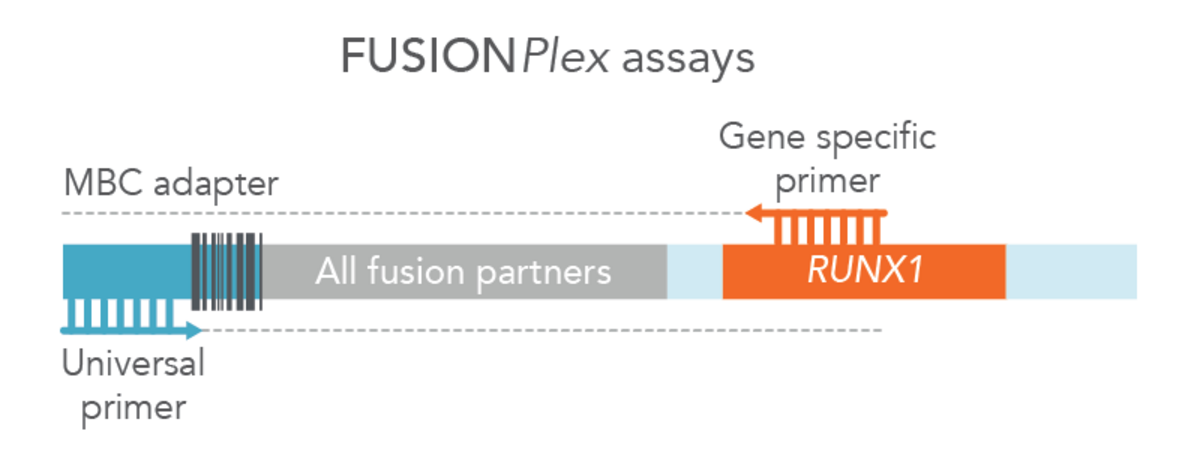
Archer blood cancer assays amplify through GC-rich regions, such as those in CEBPA, with high target coverage. Statistically significant and confident variant detection is powered by Archer’s unique outlier identification algorithm informed by normalized data to characterize position specific noise stemming from PCR and sequencing.
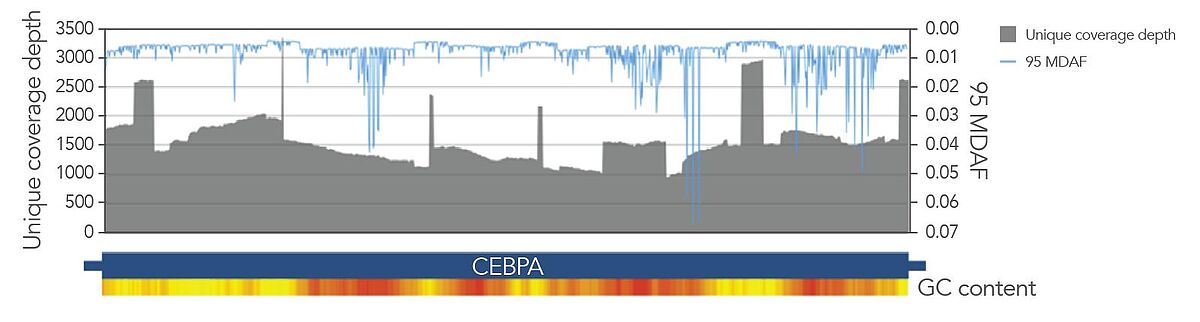
VARIANTPlex blood cancer assays also provide comprehensive detection of ITDs. AMP chemistry enables tiling across both strands of DNA with primers that amplify independently of each. When reads from these primers are analyzed with the Archer Analysis pipeline, long consensus reads are generated based on sequence similarity and aligned against the genome to identify duplication events, allowing for confident detection of ITDs, including FLT3-ITD (Table 1).
Table 1. FLT3-ITDs called at ultralow allele frequencies using the VARIANTPlex Core Myeloid panel.
A FLT3-ITD-positive cell line was titrated into a background control sample. Triplicate libraries were prepared using 200 ng total input DNA and the VARIANTPlex Core Myeloid panel. ITD calling was performed using Archer Analysis.
| FLT3-ITD fraction | Input ITD molecules* | Expected ITD called |
|---|---|---|
| 0.0 | 0 | 0/3 |
| 0.75 | 45000 | 3/3 |
| 0.075 | 4500 | 3/3 |
| 0.0075 | 450 | 3/3 |
| 0.00075 | 45 | 3/3 |
| 0.00025 | 15 | 2/3 |
*Assuming 300 haploid copies per ng.
Blood cancer panels and reagent formats fit for your lab’s needs
| Genes targeted | Genomic alterations detected | Recommended reads | ||
|---|---|---|---|---|
| DNA input | VARIANTPlex | |||
| Myeloid | 75 | SNVs, indels, ITDs | 4 M | |
| Lymphoma | 51 | SNVs, indels, CNVs | 5 M | |
| Core Myeloid | 37 | SNVs, indels, ITDs | 3 M | |
| MPN Focus | 12 | SNVs, indels, ITDs | 800 K | |
| AML Focus | 11 | SNVs, indels, ITDs | 800 K | |
| RNA input | FUSIONPlex | |||
| Pan-Heme | 199 | Fusions, SNVs, indels, expression levels | 4.5 M | |
| Heme v2 | 87 | Fusions, SNVs, indels, expression levels | 1.5 M | |
| Myeloid | 84 | Fusions, SNVs, indels, expression levels | 1.5 M | |
| Lymphoma | 125 | Fusions, SNVs, indels, expression levels | 2 M | |
| Acute Lymphoblastic Leukemia (ALL) | 81 | Fusions, SNVs, indels, expression levels | 1.5 M | |
In addition to detection of the above alterations, Archer IMMUNOVerse™ assays can provide T- and B-cell clonality, minimal residual disease (MRD), and somatic hypermutation information relevant for heme malignancies. IMMUNOVerse panels have parallel workflows with VARIANTPlex and FUSIONPlex assays, allowing for comprehensive, streamlined blood cancer profiling.
Pair DNA- and RNA-based assays to efficiently generate a complete biomarker profile
| Paired panels | Genes | Reads | Samples† per NextSeq 500/550 high output kits v2.5 |
|---|---|---|---|
| FUSIONPlex Pan-Heme | 199 | 4.5 M | 47 |
| VARIANTPlex Myeloid | 75 | 4 M | |
| FUSIONPlex Lymphoma | 125 | 2 M | 66 |
| VARIANTPlex Lymphoma | 51 | 5 M | |
| FUSIONPlex Myeloid | 84 | 1.5 M | 88 |
| VARIANTPlex Core Myeloid | 37 | 3 M |
Data assumes Archer libraries are prepared using the liquid reagent workflow.
†Sample = 1 VARIANTPlex library + 1 FUSIONPlex library
VARIANTPlex and FUSIONPlex panels can be paired for cost-effective solutions to provide SNVs, indels, ITDs, fusions, and expression alterations from a single sample.
Choice of assay reagent formats are available to suit your lab’s sample volumes.
- For high-throughput labs, liquid reagents are ideal for automated liquid handling workflows. Liquid adapters are available in a 96‑well plate format and include an increased number of dual indexes (over 36,000 unique combinations of sequencing indices) to support high-throughput sequencing.
- For manual workflow labs, single‑use, reaction‑size lyophilized reagents allow your lab to avoid potential contamination during manual pipetting and provide extended shelf life.
Both reagent formats have demonstrated excellent performance, with detection of 100% of expected variants in reference materials‡. Regardless of reagent format, all reagents are color‑coded to match protocol steps, making benchwork easy (Figure 4).
‡Data available upon request.
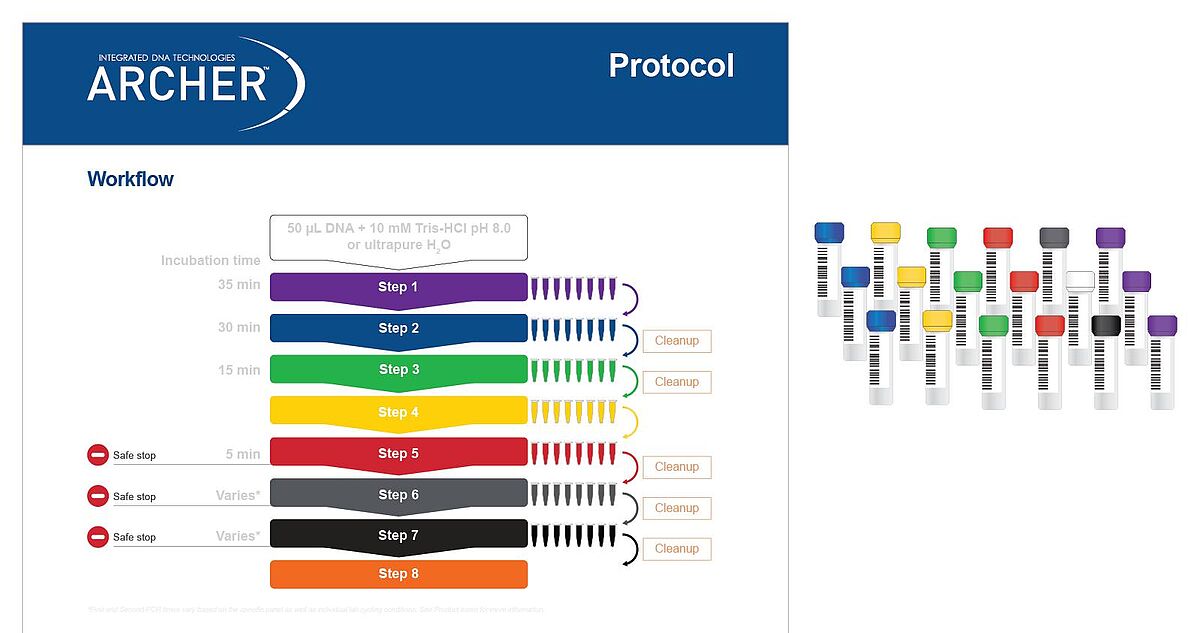
References
- Stengel A, Shahswar R, Haferlach T, et al. Whole transcriptiome detects a large number of novel fusion transcripts in patients with AML and MDS. Blood Adv. 2020. 4(21): 5393-5401.
Supplier

IDT - Integrated DNA Technologies
With over 30 years experience as a manufacturer, IDT offers innovative tools for NGS, CRISPR, qPCR and PCR. IDT offers superior quality DNA and RNA oligos, genes, gene fragments, Cas nucleases and more, with fast turnaround times!
