NGS library prep kits, panels & adapters
We offer DNA library prep kits and reagents for a wide variety of applications. Browse our selection below and contact us if you have any questions!
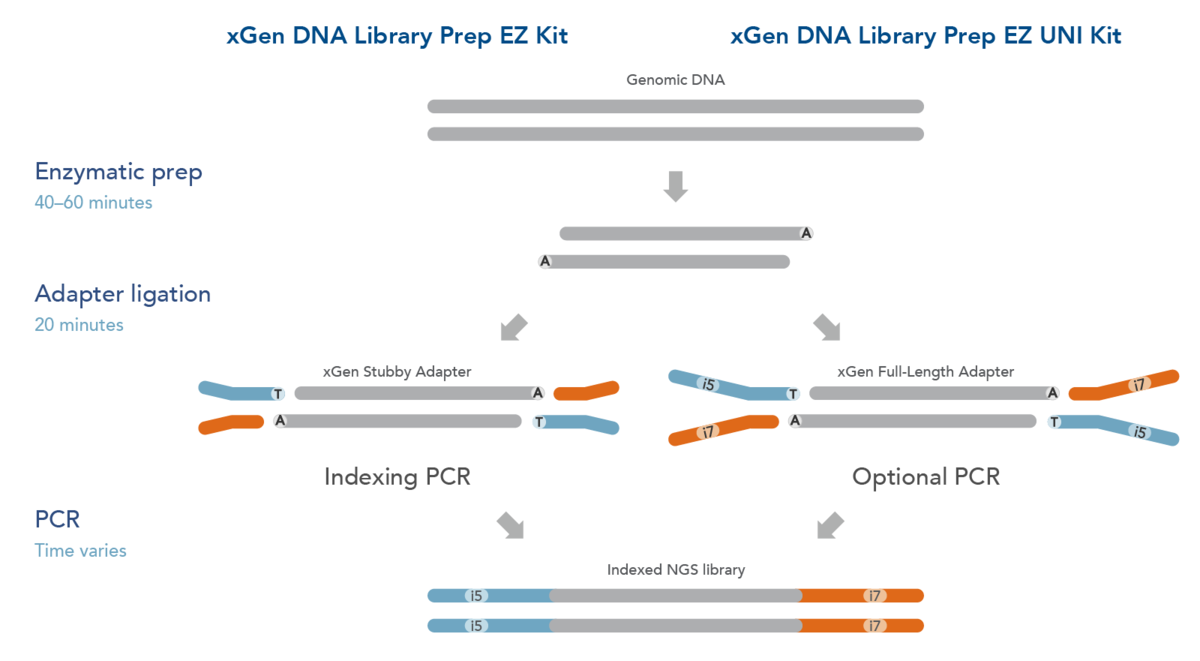
xGen™ DNA Library Prep EZ
The Easiest NGS Workflow for Routine Sequencing
xGen™ DNA Library Prep EZ are a fast, efficient and cost-effective library prep solution. Leveraging a robust enzymatic fragmentation prep and flexible indexing, you can prepare high-quality whole genome and exome libraries using a broad range of input amounts and sample types.
- High-throughput: Quick 2-hour workflow, 768-indexing, and robust fragmentation.
- Automation ready: Scripts available for Beckman, Hamilton and PerkinElmer. Fast-track help available to assist with automating on your platform.
- Budget friendly: >30% per sample over similar products on the market
xGen™ DNA Library Prep EZ Kits come in two configurations:
xGen™ DNA Library Prep EZ, All-in-one for quick implementation
Whether discovering or screening germline mutations, xGen™ DNA Library Prep EZ makes library prep more accessible to any laboratory. This kit configuration includes library prep reagents and IDT adapters compatible with several indexing options such a Single, Combinatorial Dual (up to 768-plex) or Unique Dual Indexing.
xGen™ DNA Library Prep EZ UNI: Broaden the range. Expand your scale.
Need a powerful pipeline to detect any variant across any sample quality and quantity? Combine our xGen™ DNA Library Prep EZ UNI with your choice of full-length indexed adapters (e.g. Illumina TruSeq DNA Single Indexes and TruSeq DNA CD, etc..) to create a fast, efficient, and scalable workflow to detect both rare and common variants in any size project.
“Wanted to add, I am really impressed with the uniformity in the library size! Haven’t seen that ever with a transposase based library kit.”-From a Client in Maryland
“We spoke a few weeks ago on a tech support call regarding using the Swift 2S Turbo Library Prep Kit on a mixed microbial plasmid DNA input. I wanted to let you know we followed the 10ng input parameters and it worked beautifully! Our final libraries were slightly large (~450-500bp) but still gave us high-quality sequencing reads and great data. Thank you for your help!”-Elinne Becket, Ph.D., Assistant Professor-Department of Biological Sciences, California State University, San Marcos
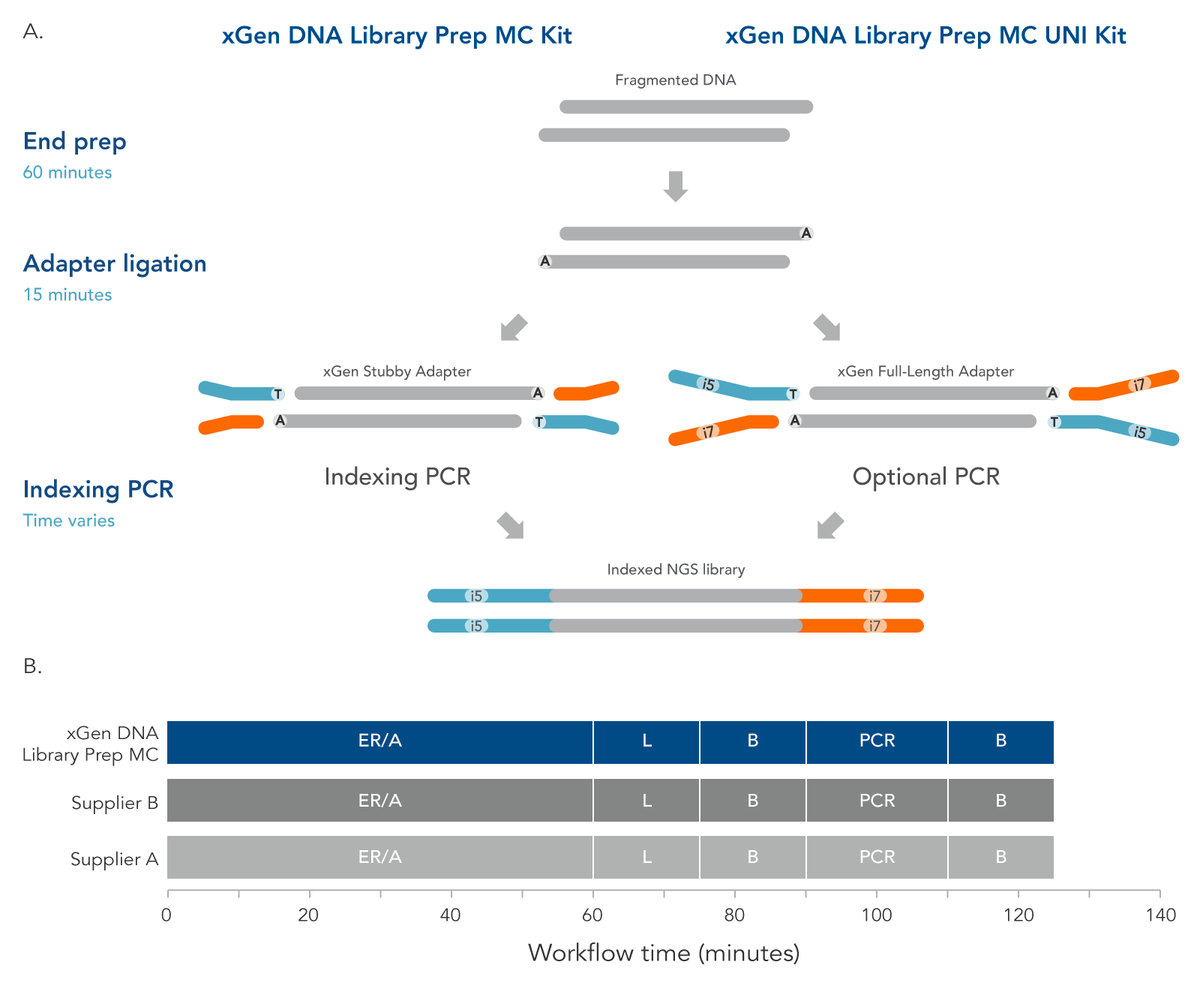
xGen™ DNA MC library preparation kits
High-throughput library prep for fragmented dsDNA - 2-hour workflow, 2 configurations
The xGen™ DNA MC Library Prep Kits offer a versatile solution that streamlines NGS sample preparation for double-stranded DNA on Illumina® sequencing platforms.
xGen™ DNA MC library prep kits come in two different workflow configurations:
xGen™ DNA library prep MC: for indexing by PCR
This kit configuration includes library prep reagents and a truncated Y adapter for compatibility with IDT xGenTM Indexing Primer kits including Single, Combinatorial Dual (CD; up to 768-plex), and Unique Dual (UD; up to 384-plex) Indexing options. This fast and efficient workflow is compatible with Normalase CD and UD Indexing Primer kits to further expedite sample processing.
xGen™ DNA library prep MC UNI: for indexing by ligation
For this kit configuration, combine our library prep kit with your choice of full-length indexed Y adapters (available from Illumina and other vendors) to create a fast and efficient PCR-free workflow. Or combine with Normalase and library amplification to further expedite sample processing.
Supported applications
- Whole genome sequencing (WGS)
- Hybridization capture of targeted genomic exome regions
- Metagenomic sequencing
- Detection of germline & somatic variant SNVs and Indels
- Copy number variation detection
Key features
- Compatible with Covaris® sheared DNA
- Supports 1 ng – 1 μg input DNA
- PCR-free libraries from 50 ng input
- Delivers data equal to Kappa and NEB kits at lower price
- Up to 768 combinatorial dual and 384 unique dual indexing
xGen™ Methyl-Seq library preparation kits
Low-input Methylation Sequencing
xGen™ Methyl-Seq library prep kit utilizes Adaptase™ technology that efficiently captures bisulfite-converted ssDNA molecules into library molecules for epigenetic research studies. The resultant libraries represent uniform and comprehensive genome coverage. The resultant libraries accurately represent sample base composition of the entire genome for the analyzed organism. The technology has quickly become the gold standard for providing the most comprehensive coverage of the methylome. It is an excellent choice for whole genome bisulfite sequencing as well as targeted bisulfite sequencing applications, such as RRBS and hybridization capture using the NimbleGen™ SeqCap™ Epi Enrichment Sysem. The Accel-NGS Methyl-Seq Kit is also compatible with bisulfite-converted DNA samples enriched by ChIP or other methods. This library preparation technology contains a uracil tolerant polymerase which is ideal for sequencing ancient DNA samples that may contain uracil nucleotides as a result of damage.
xGen™ Methyl-Seq library preparation products
- xGen™ Methyl-Seq DNA Library Kit
- The gold standard for single-base resolution of methylomes
- xGen™ Adaptase™ Module
- Prepare Single-Cell Methyl-Seq Libraries

xGen™ ssDNA & Low-input DNA library preparation kit
NGS DNA library construction from samples with limiting amounts of degraded DNA
The xGen™ ssDNA & Low-input DNA library prep kits enable users to make libraries from the impossible. Libraries can be made from precious, damaged and degraded samples by using the Adaptase™ technology. Unlike other library methods, the Adaptase technology can generate library molecules from single-stranded DNA fragments, which allow researchers to recover more of their input DNA from difficult and rare samples compared to other commercially-available products-for Ilumina® NGS Platforms
The xGen™ ssDNA & Low-input DNA library prep kits will yield libraries in 2 hours from as little as 10 pg of input and fragments as short as 40 bp. The kits work with precious and difficult-to-use samples, such as single-stranded DNA, heat-denatured pathogenic samples (microbial DNA), viral DNA, first-strand cDNA, ancient DNA, ChIP-Seq, and degraded DNA from FFPE samples. Accel-NGS 1S Library Kits are the best choice for users needing to sequence treasured or difficult-to-process samples, which cannot be sequenced by other methods. Expand your research by processing precious, damaged and degraded samples on either the Illumina® or Ion Torrent™ NGS platforms.
xGen™ ssDNA & low-input DNA library preparation products
- xGen™ ssDNA & low-input DNA library prep kits
- DNA Library Preparation of Difficult Samples
- Rescue valuable sequencing data from precious samples
“We choose this kit for construction of sequencing libraries when available sample is low, the quality is uncertain, or when we want to capture single stranded as well as double stranded DNA. We have found the kit easy to use with well labeled components and a straightforward protocol. We find that libraries rarely fail with this kit even when starting material is highly degraded. The availability of unique dual indexes is also key to allow sequencing on any available Illumina platform. With regards the samples from the NASA Twins study, we were most interested in being able to detect single-stranded DNA viruses as well as all other organisms as part of our shotgun metagenome sequencing project, and this drove the decision to use this kit.”
Stefan J Green, Ph.D. Director, Sequencing Core, Associate Director, RRC and Kevin Kunstman, MS Assistant Director Sequencing Core, University of Illinois at Chicago
“I would like to give a summary that we tried to use a competitor’s kit first for our project but it ended up that we needed to do 17 PCR cycles to achieve the concentration of libraries that we needed. But what that meant was that there would be increased duplication rate when we did next-generation sequencing. With your kit we only needed to do 4-5 PCR cycles and our duplication rate was 7-13% when using HiSeq4000 for deep sequencing. We were interested in both ssDNA and dsDNA. We are happy with your product.”
Premi Haynes Ph.D., Senior Fellow in the Laboratory of Dr. Daniel Miller, University of Washington
xGen™ cfDNA & FFPE DNA Library Preparation Kit
High library complexity from low quality samples
The xGen cfDNA & FFPE DNA Library Preparation Kit empowers you with accurate variant identification from degraded and low-input research samples.
- Get high conversion rates compared to TA-ligation-based methods with novel ligase and highly modified adapters
- Identify variants at ≥1% variant allele frequency (VAF)
- Get data from even highly degraded research samples
NxSeq® AmpFREE low DNA library kits
The highest efficiency, low input, pcr-free fragment library prep kit available at the lowest cost!
- Low Input: Requires as little as 75 ng of sheared input DNA allowing use of limiting samples.
- High Efficiency: Optimized adaptor ligationproduces more sequenceable fragments in each library, yielding better coverage & depth from single or multiplexed libraries.
- PCR-free: Prevents the introduction of PCR-bias, providing more uniform coverage.
- Fast: 2 hour, 10 minute protocol saves you time and gets your samples on the sequencer sooner.
- Affordable: Best priced and best performing kit available.
NxSeq® UltraLow DNA library kit
Lowest input, highest efficiency, Illumina-compatible DNA fragment library prep kit
- High Quality Data: High efficiency adaptor ligation produces complex libraries that yield improved sequencing depth uniformity and better coverage with fewer zero coverage regions
- Sensitive: Construct DNA fragment libraries from as little as 50 pg to as much 75 ng of sheared/fragmented DNA
- Minimal Bias: Robust, uniform PCR amplification improves coverage uniformity when working with low input amounts of genomic DNA
- Flexible:Extensively tested in de novo whole genome sequencing or resequencing, but compatible with other applications such as exome-seq, ChIP-seq and FFPE DNA samples.
- Fast: 3 hour protocol gets your samples on the sequencer quicker
- High Value: Cost-effective library and indexing kits which produce excellent sequencing results
For research use only. Not for use in diagnostic procedures. Unless otherwise agreed to in writing, IDT does not intend for these products to be used in clinical applications and does not warrant their fitness or suitability for any clinical diagnostic use. Purchaser is solely responsible for all decisions regarding the use of these products and any associated regulatory or legal obligations.